Therapeutic Antibody Design – The AI-assisted Future Of Lead Optimization
Nov 16, 2022
In our August 2022 manuscript published in bioRxiv, we explored using our machine learning models to simultaneously optimize binding affinity and naturalness scores of the therapeutic antibody trastuzumab. Read it to learn more about how we are leveraging generative AI and scalable biological data to lead the protein drug creation revolution:
The therapeutic antibody field has experienced explosive growth as biologics have become the predominant class of new drugs developed by major players in the pharmaceutical industry. Antibodies are natural drugs produced by our immune system and very promising therapeutics, especially in cancer and immune diseases. Their power comes from natural selectivity – they can bind to specific target proteins, which can be engineered to block or change biological processes underlying disease. Also, a properly designed therapeutic antibody will be recognized by our body as part of natural defenses without invoking an immune response or unwanted side effects.
Over the past several years, biologics progressed toward being the top best-selling drugs – seven out of ten blockbusters in 2022 – creating an extremely competitive therapeutic niche. Identifying novel methods to optimize these drug candidates’ safety, efficacy, and manufacturability in the shortest time possible has become more critical than ever.
How To Develop A Therapeutic Antibody?
Therapeutic antibodies traditionally undergo five significant sequential stages leading up to preclinical development. First, the drug’s target is investigated for its relevance in the disease based on primary research. Then the antibody is generated by immunizing an animal and growing its immune cells that produce the antibodies. These antibodies are initially tested in the lab, and those that look most promising pass on to the next step, initial lead selection. These lead drug candidates undergo lead optimization, a critical stage in which the antibodies are honed for biological activity and properties that will impact the drug’s efficacy, safety, and developability. Finally, the optimized lead candidates go through preclinical enabling to make them suitable for animal testing.
Lead optimization is one of the most vital steps in antibody development. It plays an essential role in maximizing an antibody’s likelihood of clinical success and ultimately becoming an approved drug. Unfortunately, this step is also extremely expensive, time-consuming, and plagued with a high rate of failure. One of the biggest challenges is that lead optimization must simultaneously assure efficacy, safety, and developability. It is a very common case that perfecting one parameter ruins another – a process that has often been described as playing a game of ‘whack-a-mole’ where improving one property may lead to unintended effects on other properties and so on in a perpetual cycle that may never yield an ideal candidate.
Two of the most important properties to optimize – or moles to simultaneously knock down – before preclinical studies are efficacy and safety. These are indicators that an antibody will have a strong therapeutic effect and be less likely to have severe side effects. In order to have high efficacy a good therapeutic antibody candidate will ideally only attach, or bind, to very specific cells in your body, such as cancer cells. This allows the cancer cell to be destroyed by the immune system while leaving healthy surrounding tissue intact. The antibody binding must be not only specific, but also very strong and long-lasting to give immune cells time to kill the cancer cells. The strength of this binding is defined as affinity, and is a key metric drug developers seek to optimize for therapeutic antibodies.
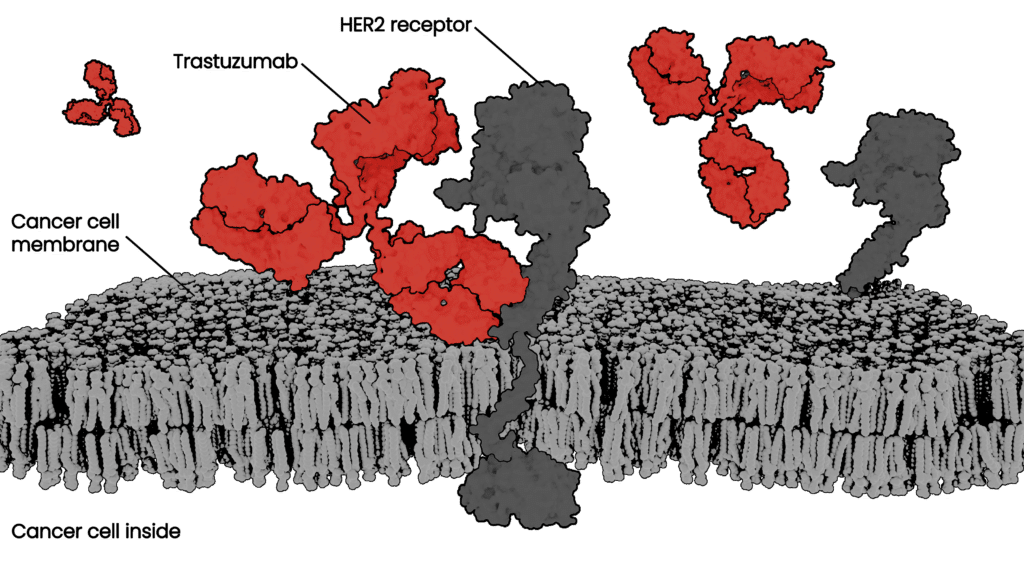
Another indicator of whether a therapeutic antibody will produce severe side effects is immunogenicity – or whether the antibody is likely to trigger an unwanted immune response. Immunogenicity in particular, can be a tricky parameter to gauge in a single experiment due to the complexity of the immune system and all the different cells and proteins a therapeutic antibody may encounter on the way to its target. However, it has been hypothesized that the more similar an antibody is to those occurring in nature, the less likely they will be to create an immune response. With this in mind, we have been developing a new metric called “naturalness”. We believe that therapeutic antibodies with high naturalness scores will be less likely to trigger an immune response in patients, leading to fewer adverse effects and better outcomes.
Stronger, Better, Faster: Machine Learning To The Rescue
To design a promising lead drug candidate for further development, you need to find antibodies with several different properties, all falling within a small range. It’s like searching for a needle in a haystack, except the haystack is larger than the known universe, and the needle is smaller than a speck of sand. Using traditional discovery methods, this can be a long, expensive process with a little probability of success. At Absci we’ve designed our generative AI models to significantly speed up and improve the lead optimization process compared to traditional methods that rely solely on wet lab screening. Using our proprietary wet lab assays we generate the massive amounts of biological data needed to enable the predictive power of machine learning. Using computers to design, test, and run experiments in silico (on the computer rather than at a lab bench) we can dramatically speed up the lead optimization process and discover unique, generative variants that could never be found using standard methods.
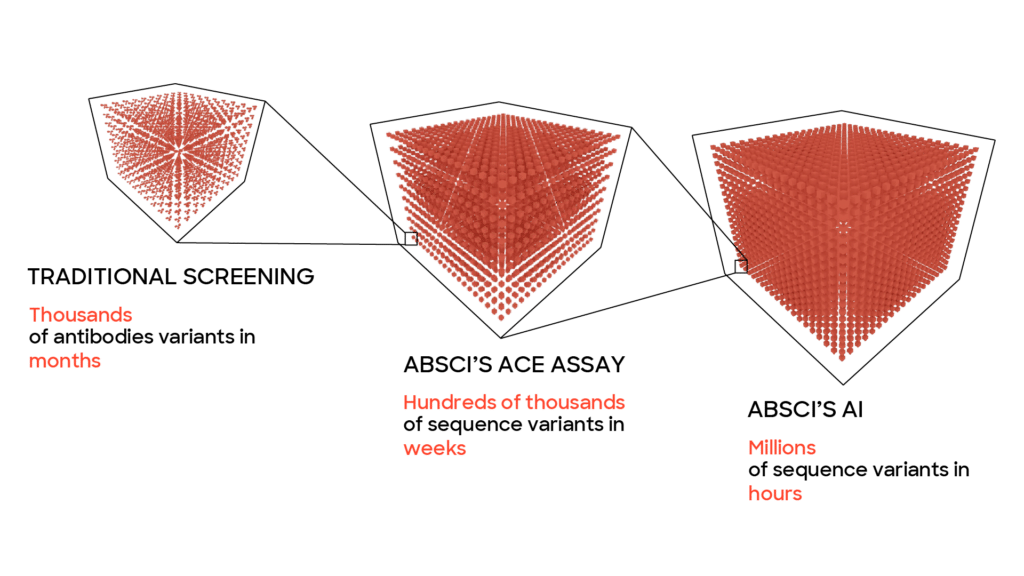
One reason this method is able to work faster than traditional methods is that machine learning models allow us to test a much larger number of potential drugs – far more than would be possible or practical to explore exclusively in a wet lab due to time and money considerations. With our platform, we can quickly probe a wide variety of vastly different potential drugs and then hone in on the most promising ones extremely quickly. To do what would take years using traditional methods can be done in months or weeks. For the lead optimization phase of a project this enables the computer to play our metaphorical whack-a-mole game at a breakneck pace until it finds an antibody where all the parameters have been optimized and the therapeutic antibody can continue to preclinical studies.
Let’s Work Together to Bring Therapeutic Antibodies to Patients Faster!
We work with our partners on target and antibody discovery, AI-assisted antibody optimization, and site-specific payload conjugation via nsAA incorporation. Connect with us today to see how we can help you create better biologics.