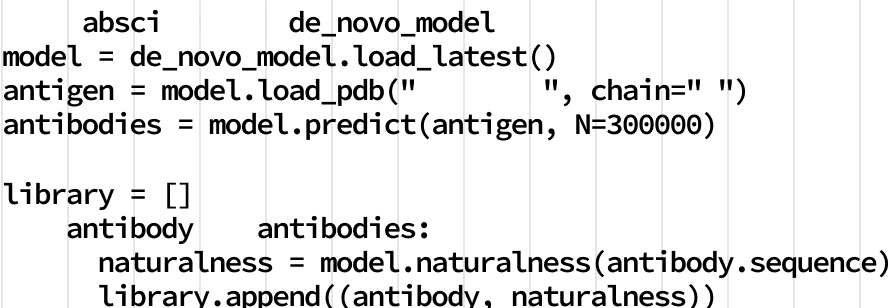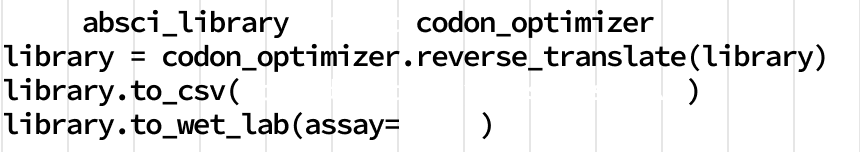
Absci Technology
A generative AI drug discovery platform for new medicines.


Our Process
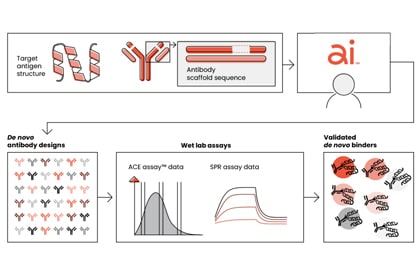
AI promises to revolutionize drug discovery, but advances in drug creation also depend on scalable wet lab technologies to produce and validate biological data at scale. We've built the team and technology to realize this opportunity.
Our Integrated Drug Creation™ platform combines the data to train, the AI to create, and the wet lab to validate millions of AI-generated designs a week. It can take us from AI-designed antibodies to wet lab-validated candidates in as little as six weeks.
Data to Train
Proprietary wet-lab assays generating billions of protein-protein interaction data to train ML models.
AI to Create
Generative AI engine to create novel designs of antibodies/next-gen biologics.
Wet Lab
to Validate
Scalable wet lab infrastructure operationalized to validate AI generated designs in a matter of weeks.
Data to Train
We begin with data.
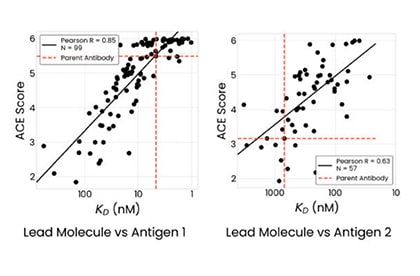
Our SoluPro® technology is a multiplex synthetic biology approach that overcomes the limitations of today's highest-throughput automation labs. It constructs billions of genetically-distinct cells, each containing instructions to make one version of a protein of interest, as well as a different assortment of folding and expression solutions. Our ACE Assay then evaluates and sorts hundreds of millions of designs to collect the best hits - versions of the protein-of-interest, based on target binding, protein quality, and expression titer. We use this high-quality, high-throughput data to train our AI models.
AI to Create
Generative AI expands the solution space.
Most drug companies use classical discovery techniques to search existing antibody libraries to develop drug candidates with incremental improvements. At Absci, our drug discovery platform can design new therapeutics using the same kind of AI that is renowned for creating text and images from natural language prompts. Working from massive biology datasets, we apply generative AI to design optimal drug candidates based on target affinity, safety, manufacturability, and other traits. This enables us to create completely new antibody designs in the computer - a true breakthrough that ushers in a new era of in silico drug design.
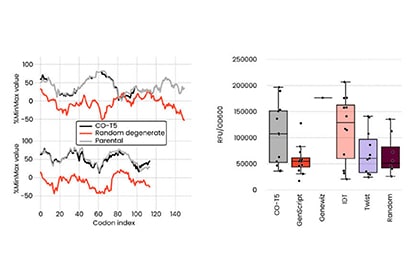
WET Lab to Validate
AI models that are proven in the lab.
We support our generative AI designs with our wet lab's massive-throughput functional validation capabilities. It can take us from AI-designed antibodies to wet lab-validated candidates in as little as six weeks. The quality and scale of wet lab data give us incredible training data, propelling our iterative design-build-test-learn cycle.

How can our Integrated Drug Creation™ platform help you go from drug discovery to drug creation?