Shaheed Abdulhaqq on the holy grail of HIV research – and the path that brought him full circle in the fight against the virus.
More than 40 years after the AIDS pandemic began, there is no vaccine or cure for HIV. Antiretroviral therapies (ARTs) help many people live longer, healthier lives, but they do not completely eliminate the virus and must be taken for life. Additionally, the cost and inaccessibility of these drugs disproportionately affect millions of people from low-income and marginalized communities.
“HIV is a challenging virus,” says Shaheed Abdulhaqq, Absci Group Leader and HIV researcher. “It mutates quickly and attacks one of the most important types of immune cells in the body,” he says, “so the very thing that fights HIV is under attack.”
A new collaboration between Caltech and Absci aims to change that. A grant to Caltech from the Bill & Melinda Gates Foundation is bringing together state-of-the-art biotech and research expertise in fields like immunology, protein design, and generative AI with the ambitious goal of creating a better, more accessible HIV therapeutic vaccination – one that both treats and protects against infection from multiple HIV-1 strains.
Using generative AI to unlock HIV
As part of the team, Shaheed is eager to apply generative AI to unlock HIV, but the challenge is clear. HIV evades treatment by an ever-changing variety of proteins on its surface, making it impossible so far for researchers to design a drug that can find and target it. Shaheed says the key to defeating HIV may be in the way it infects cells.
When HIV comes in contact with a human CD4+ T cell, he says, “it opens up like a flower.” This exposes the parts of the virus that infect the human immune cell. But unlike the virus surface, this exposed portion is conserved and unchanging, representing a promising drug target. Caltech researchers propose creating a novel HIV therapy that first exposes and then binds to a conserved site on HIV-1. Absci’s role in the project is to design the antibody or antibody-like protein to bind to the exposed region and disable the virus.
“We’re trying to target the virus in a way that no one else has ever done before,” says Shaheed. “If we are successful, this could be the holy grail the HIV field has been seeking for decades.”
Academic roots coming full circle
For Shaheed, this new Gates Foundation project feels like coming full circle. As a research assistant professor at Oregon Health & Science University, Shaheed worked on an HIV project supported by the foundation, and he even won a foundation award for some of his work.
“The Gates Foundation is a wonderful organization, and I don’t know where the field of HIV research would be without it,” Shaheed says.
He began working on HIV as a graduate student at the University of Pennsylvania, where he was interested in understanding why some people seemed impervious to HIV infection.
“There are many routes to infection —intravenous drug use, sexual intercourse with an HIV partner, or from mother to unborn child. But for whatever reason, some people just don’t become infected. And my research at that stage was really focused on: what is special about these people? Why are they not getting infected?”
Landing in the Pacific Northwest
His graduate work earned him offers in Houston and Portland, though in the end he accepted a postdoc position at Oregon Health & Science University for reasons unrelated to research.
“I remember flying into Houston and looking down at this humongous, sprawling city. There was concrete and buildings everywhere, with maybe a speck of a tree here or there.” He compares that with flying into Portland. “You come in over Mt. Hood, with the Columbia River off to one side, and then the Willamette Valley reveals itself, and it’s just filled with trees. It’s utterly beautiful and breathtaking — I knew I wanted to live here.”
The research was also great. In Jonah Sacha’s lab, Shaheed worked on an HIV vaccine using cytomegalovirus (CMV). The vaccine was somewhat unique in that it didn’t prevent infection, but it induced a strong immune response that ultimately clears HIV completely — akin to some of the super-responders he studied earlier.
“It basically controls the infection and then completely clears it. Very unique. No other vaccine I’ve ever seen works in quite that way.”
Biotech life in the Pacific Northwest
In the five years working on that project, Shaheed had moved up to research assistant professor and had a couple of grants under his belt. Then the pandemic came, and his lab was effectively shut down. It prompted Shaheed to rethink his career.
“I could stay on the academic track,” he says, “which meant a lot of grant writing, and that’s a really hard endeavor and very far away from the research.” Shaheed talked with a lot of grad school friends who went into big pharma and biotech startups. “They seemed happy and passionate about the impacts they were making, and they were a lot closer to clinical impact than I could have been in academia.”
In addition to seeking impactful work, he knew he didn’t want to leave the area. That was about the time that one of his friends mentioned a cool Vancouver biotech firm that was looking for someone with flow cytometry experience. It sounded like a good match for someone like Shaheed — flow cytometry is a key tool in cellular immunology. He met with the Absci team, and when the offer came, Shaheed jumped at it.
“I fell in love with the environment,” he recalls. “I don’t know that we were calling ourselves Unlimiters back then, but the Unlimiter spirit was really strong and alive. I didn’t care about any other interviews. Absci was where I wanted to be.”

Building a wet lab to enable AI
For ten years, flow cytometry was Shaheed’s tool of choice for finding potential HIV vaccines – like finding a needle in a haystack. Now, Absci was looking to Shaheed for ways to improve the use of its proprietary ACE Assay™ technology – developed by Absci scientist Jia Liu – as a high-throughput screening tool for E. coli, its workhorse microbe. Flow cytometry is typically used with large mammalian cells, which are hundreds of times bigger than E. coli. Were there ways to improve the ACE Assay workflow?
“It was a humbling experience,” Shaheed says. “The data we were looking for was in this space that I typically thought of as debris. When studying mammalian cells, you ignore it because it’s fragments, contaminants, and junk you just don’t care about. But with E. coli, all of our signal was in that area.”
Shaheed and the team set about developing the protocols to filter the signal from the noise at a whole new level, including new assays, improving sample preparation, and changes to the instrument itself. It also demanded a more innovative approach, with the team asking itself: What unique features of E. coli can we leverage to use flow cytometry to analyze and assess bacteria?
Shaheed isn’t giving away any secrets, but he says it was great fun — and a privilege — to plow into the problem, mentor the team, and help guide novel research and development of one of Absci’s core technologies – which in effect makes the impossible possible.
Today, Shaheed leads Absci’s ACE Assay team. What the ACE Assay platform can do in the lab is central to what Absci can do with AI. Shaheed says the ACE Assay technology is perfect for screening millions of AI-designed antibodies to determine which ones bind to a target. In addition, all that data about what binds and what doesn’t creates a huge volume of training data for our AI models.
“That enables our models to define a binder that’s even tighter than anything that was in the training set,” Shaheed says.
Believing in the impossible
As the Caltech project gets underway, Shaheed again feels the same sense of awe at the challenge ahead. To be successful, it will demand innovation and collaboration across multiple different teams: AI, Strain Engineering, Biophysics, Protein Engineering, Strain Engineering, Sequencing Development, and ACE Assay Teams.
“It’s a very challenging project,” but Shaheed says that’s not a discouragement. “At Absci, we’re frequently doing things that no one else has ever done before.” This project is no different.
“Absci is a generative AI drug creation company, but we also have a passion for solving really difficult problems to improve the world,” says Shaheed. “It’s a beautiful merger of creating products to help patients and also enjoying the adventure of cutting-edge research. It’s that beautiful synthesis that really makes me appreciate Absci and our work.”
Harnessing AI for biological innovation

Absci Founder & CEO Sean McClain spoke at the Senate AI Insights Forum on October 24, 2023, on maintaining U.S. leadership in AI, minimizing risks, and harnessing AI for the greatest good.
By Sean McClain, Absci Founder & CEO
The transformative power of artificial intelligence (AI) is indisputable, with its influence resonating across diverse sectors. However, the conversation around AI often focuses on mainstream applications like ChatGPT and DALL-E, with less attention paid to the transformative potential of AI in healthcare and biotech. At Absci, we are using AI with the goal of accelerating drug discovery to create better medicines for everyone and respond rapidly to health crises and outbreaks. That stands to impact people’s daily lives in a profound way.
To share this perspective, I was invited to the second Senate AI Insights Forum hosted by Senate Majority Leader Chuck Schumer and Senators Rounds, Heinrich, and Young. This event, attended by influential figures such as Marc Andreessen and John Doerr, and leading academics and civil society leaders, aimed to foster discussion on how the U.S. can sustain its leading position in AI while ensuring safety. Here are some of the critical points I shared with lawmakers.
Data is paramount
The role of data in AI for biology cannot be overstated. Quality data is the backbone of any successful AI application. While public data repositories lay the groundwork, proprietary high-throughput biological data is the key to unlocking AI’s potential in drug discovery. Unlike ChatGPT or DALL-E, which have the entire internet as training data, biological data is dwarfed in comparison and is the bottleneck to scaling AI in biology. Only by scaling high-quality biological data via platforms such as Absci’s can our nation win the race in AI for biology.
Data is the critical next step to unlock AI’s potential in biology. The U.S. must sustain its leadership in AI by accelerating investments, particularly in producing and protecting high-quality biological data. This is essential for timely and effective responses to emerging health crises.
Several public data sources are vital to U.S. innovation in biotech and beyond. These open-source datasets are essential resources for drug discovery – we should support them in every way we can through policies that encourage collaborative scientific advancement while instituting robust safeguards against potential bioterrorism and unintentional accidents.
At Absci, we also realize that open-source data alone cannot scale for AI. That is why we build on public data sources by adding our own experimental wet lab data. Our proprietary synthetic biology platform generates the massive amounts of biological data needed to enable the predictive power of machine learning. Then, we use that data to design and run experiments in silico (on the computer rather than at a lab bench). This dramatically speeds up the discovery process at a scale and pace that could never be applied using standard methods. Supporting the development of a vibrant biotechnology community with pooled proprietary datasets will help set the U.S. apart when scaling biological data for AI
Safeguarding against the weaponization of AI
As much as AI in biology can be used to transform the lives of patients for the better, without proper safeguards, this same technology in the hands of bad actors can be weaponized to do harm at unprecedented levels. Absci’s vision is to deliver breakthrough medicines at the click of a button. By the same principle, bad actors have the potential to leverage AI to create biowarfare and bioterrorism agents at the click of a button. The consequences of that would be devastating for America and the world.
The critical safeguard against the threat of bioterrorism is, once again, data. We need a U.S. strategy to create and maintain data and AI superiority that positions U.S. innovators to have superior tools at their disposal to counter terrorist organizations or other opposing forces. Policymaking should prepare for this threat not as an “if” but rather as a “when.” The government can play a key role in mitigating harms through collaborations and partnerships with and between U.S. innovators for AI rapid response programs to counter bioterrorism. If we look at Operation Warp Speed, American innovation brought three life-saving Covid vaccines to the world in record time. Let’s borrow from those winning ingredients to accelerate a U.S. era of AI leadership through focused partnerships between the government and U.S. innovators aimed at solving massive societal issues that affect us all: pandemic preparedness and protecting U.S. citizens from the threat of bioterrorism.
One-size AI policy does not fit all
The unique nature of AI in biology vis a vis the general discourse around AI calls for a nuanced approach by policymakers. Some advocate that generative AI should be heavily regulated to prevent its potential misuse. Others think it should be lightly regulated so as not to stifle innovation and public benefit. An all-or-nothing approach like this may be the wrong way to think about it. That’s because generative AI introduces different kinds of challenges to different industries.
For example, AI-designed drugs go through the same rigorous FDA approval process as drugs designed by conventional drug discovery campaigns to ensure their safety and effectiveness, so additional regulations on the AI (which in this case is a tool to discover the drug candidate) would amount to double regulating. This could result in stifling the very innovation that could improve the efficiency and success rates of getting new medicines to patients. On the other hand, we must ensure we have access to high-quality training data that is free from biases that may discriminate against underrepresented groups. Regulators could establish guidelines to ensure the quality of the underlying data used in designing AI drugs.
Regulation of AI in biology is a delicate balancing act. Policymakers must refrain from imposing overly restrictive regulations, as this could hinder innovation. Furthermore, AI-derived medications still face traditional FDA scrutiny. Therefore, regulations should focus on ensuring the integrity and neutrality of data, which are crucial for developing safe and effective AI-driven drugs.
Lead today or follow tomorrow
Generative AI might be the fastest-moving field in the history of technology. The United States must lead today, or we will be forced to follow tomorrow. The stakes are too high for the government or industry to tackle AI policy-making alone. For our part, Absci can help guide policy constructively and positively at the intersection of AI and biology. It’s also important to me to remind us all not to lose sight of the huge opportunity for patients and our society. Every minute of every day, patients wait for a cure, and our nation has a technology that could potentially deliver breakthrough therapeutics at the click of a button.
It was an immense honor to offer our perspective from Absci’s work in AI drug creation to the important and time-sensitive dialogue around AI and policymaking. I thank Leader Schumer, and Senators Rounds, Heinrich, and Young for inviting me. We look forward to continuing to share our insights in AI and biology in helping to guide generative AI toward the greatest public good.
Target-based drug discovery – looking for Holy Grails of pharmacology
Target-based drug discovery is the most common strategy for developing new medications. However, when it comes to clinical trials, most new drugs fail due to inadequate cure rate (efficacy) or safety concerns. The extremely low success rate – only about 12% of drugs make it to the market – suggests that a “wrong” target has been selected during the target discovery process.
Identifying the molecules and mechanisms primarily involved in disease is an essential first step in discovering targets that can be acted upon therapeutically. This initial step traditionally begins with a manual investigation of scientific literature and biomedical databases to gather evidence linking particular proteins to disease. This quest relies on using bioinformatics tools and collecting experimental evidence supporting the hypothesis that changing the molecule’s function will affect a patient’s health.
Drug target – what is it, exactly?
A drug target can be defined as a molecule in the body, usually a protein, that is intrinsically associated with a particular disease. A perfect drug target should be involved in a crucial biological pathway, be functionally and structurally well characterized, and be druggable – capable of binding to therapeutic molecules.
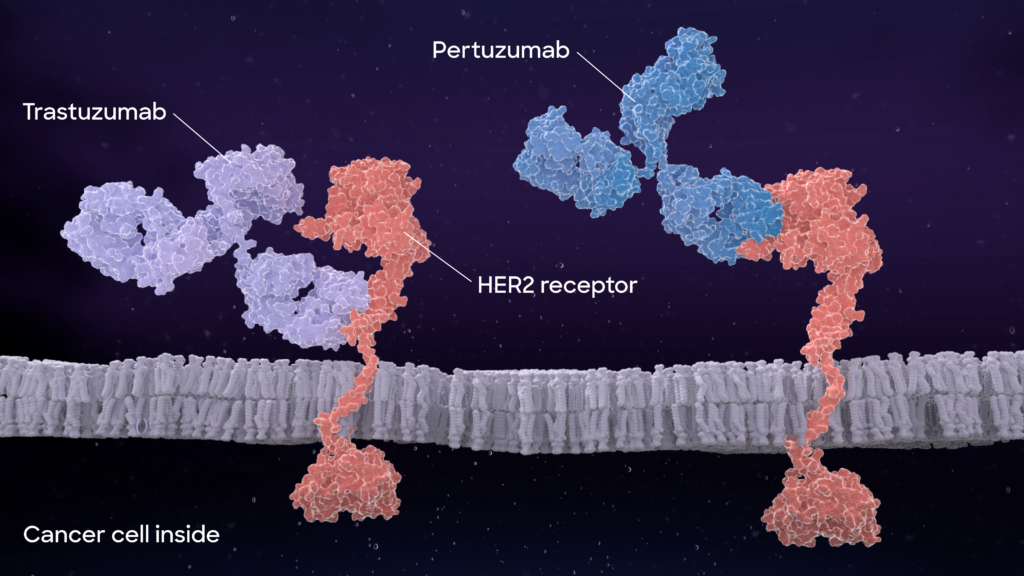
In a perfect scenario, a drug (here a therapeutic antibody) should bind only to the intended target and initiate its structural or functional change for therapeutic effect.
It sounds simple, but in reality, it is one of the most challenging ventures undertaken by humanity. The search for new pharmacological targets is often hampered by partial information about disease pathogenesis, usually consisting of conflicting data. Much knowledge about molecules in our bodies is based on indirect evidence from animal, cell, and tissue culture experiments. Unfortunately, therapies based on animal model biology often fail to replicate when tested in human trials. Moreover, many proteins are druggable based on their structure – they will bind the drug – but this will not lead to therapeutic benefit.
Researchers also rely on human epidemiological studies, for example, comparing blood levels of assumed disease-related molecules. Still, these results are subject to erroneously assigning cause and effect or misidentifying correlation for causation. Thus, there’s a desperate need for more robust information about targets and their potential clinical efficacy.
Why can’t we make more drugs?
The current approach of target-based drug design is not limited by the technical hurdles to produce the drugs but by the need for progress in identifying novel biological targets. In theory, all biological macromolecules – about an estimated 10,000 – are potentially druggable targets, but only about 400 are currently approved. Over the past 50 years, the number of drugs sharing the exact mechanism of action or target has averaged about 4, ranging from 2-14 drugs per single target. So, it seems companies struggling to identify new targets fight over those recognized as therapeutically beneficial, trying to make a better mouse trap than their competitors.
In recent years, human genomics has been increasingly revealing its power. The underlying idea is to use the powerful tool of human genetic association to identify key drug targets involved in the disease, which can be changed or blocked by drugs. However, these approaches require sifting through data from mega-repositories and large case-control consortium-type studies. The biggest challenge here is the data originating from multiple sources with unclear predictability and consistency. In addition, those associations require further studies to resolve casual relationships to confirm that the putative targets are indeed involved in disease pathogenesis and susceptible to therapeutic modulation.
Can we learn from our bodies how to pick drug targets?
Our bodies can fight and cure diseases, including cancer and infections. Could we investigate how our immune systems pick the targets to apply self-therapy? Absci is pioneering the discovery of new drug targets utilizing a process we call reverse immunology. Our approach is grounded in natural immunology, and we use our proprietary, integrated wet lab screening and deep learning to open the door to new drug target identification.
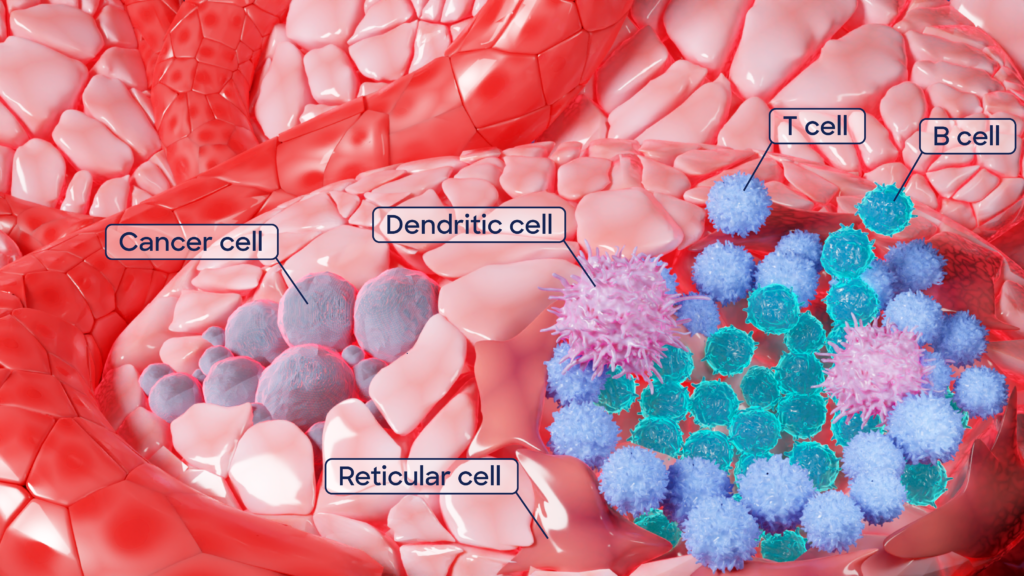
How does reverse immunology work? It starts with patient samples; these could be a patient population that displays exceptional immune responses in studies. The secret to the new targets lies in a particular tissue, called tertiary lymphoid structures, or TLSs. They resemble lymph nodes – key hubs of immune cell manufacturing – however, they don’t form in healthy tissue but only in places with chronic inflammation, infection, transplanted organs, autoimmunity, or cancer. How they develop is still under investigation, but they are most likely our body’s response to long-lasting exposure to inflammation caused by unhealthy conditions. TLSs are embedded special forces of the immune system, deployed to disease-affected tissues to produce defenses locally. A small army that identifies the enemy and delivers the ultimate weapon, target-specific antibodies, against the disease-causing cells or pathogen.
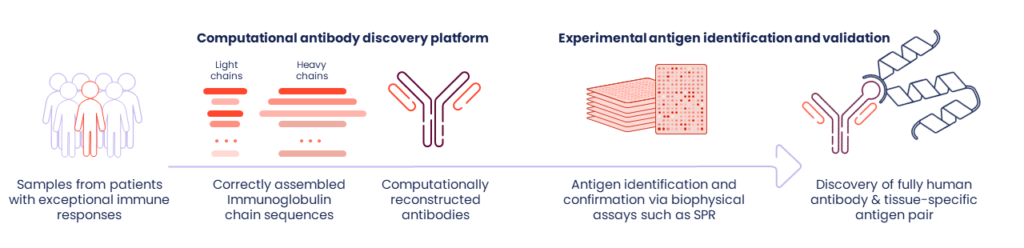
We call our approach “reverse immunology” because we start with those immune cells from TLSs and identify the antibodies they produce in the greatest quantities. What we are looking for, specifically, are the antibody blueprints from immune cells – RNA sequences, which we assemble computationally. We then use those antibody sequences to identify target antigens – rather than starting with a target and generating or fishing for an antibody. Our process is then reversed.
After reconstructing fully human antibody sequences in silico, we produce these antibodies in the lab. We use high throughput proteomic screening methods to determine their target antigens in an experimental process we call “deorphaning.” At the end of this reverse immunology process, we have the immunologic target paired with a fully human monoclonal antibody that can be used as a starting point for drug development.
Our methods are exceptionally robust – they do not require the isolation of single immune cells or the processing of fresh tumor tissues. Instead, we can work with RNA-seq data we generate from banked tissue samples, including archival specimens collected by academic consortia, clinical trials, and commercial biobanks. Thus, we can direct our technology toward curated source tissues selected for desired disease and therapeutic response profiles.
We have constructed an extensive library of antibody-target pairs. Some antigens we’ve found are the usual suspects already known to be either essential mediators of immuno-oncology responsiveness in the case of cancers or neutralizing antibodies in the case of SARS-CoV2. Importantly, we’ve also identified many novel antigens that may have promise as therapeutic targets, and we have fully human antibodies in hand.
Absci’s reverse immunology approach is powerful and efficient, offering an unbiased antibody and target discovery method. We believe it will allow us to create better biologics for patients faster.
Learn how we can work together to find new targets
In our August 2022 manuscript published in bioRxiv, we explored using our machine learning models to simultaneously optimize binding affinity and naturalness scores of the therapeutic antibody trastuzumab. Read it to learn more about how we are leveraging generative AI and scalable biological data to lead the protein drug creation revolution:
The therapeutic antibody field has experienced explosive growth as biologics have become the predominant class of new drugs developed by major players in the pharmaceutical industry. Antibodies are natural drugs produced by our immune system and very promising therapeutics, especially in cancer and immune diseases. Their power comes from natural selectivity – they can bind to specific target proteins, which can be engineered to block or change biological processes underlying disease. Also, a properly designed therapeutic antibody will be recognized by our body as part of natural defenses without invoking an immune response or unwanted side effects.
Over the past several years, biologics progressed toward being the top best-selling drugs – seven out of ten blockbusters in 2022 – creating an extremely competitive therapeutic niche. Identifying novel methods to optimize these drug candidates’ safety, efficacy, and manufacturability in the shortest time possible has become more critical than ever.
How To Develop A Therapeutic Antibody?
Therapeutic antibodies traditionally undergo five significant sequential stages leading up to preclinical development. First, the drug’s target is investigated for its relevance in the disease based on primary research. Then the antibody is generated by immunizing an animal and growing its immune cells that produce the antibodies. These antibodies are initially tested in the lab, and those that look most promising pass on to the next step, initial lead selection. These lead drug candidates undergo lead optimization, a critical stage in which the antibodies are honed for biological activity and properties that will impact the drug’s efficacy, safety, and developability. Finally, the optimized lead candidates go through preclinical enabling to make them suitable for animal testing.
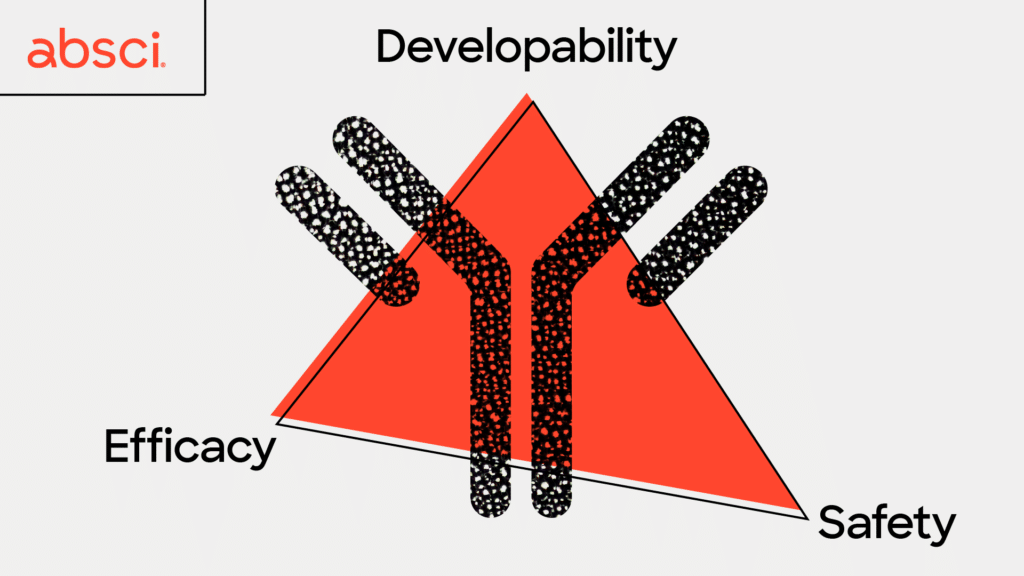
Lead optimization is one of the most vital steps in antibody development. It plays an essential role in maximizing an antibody’s likelihood of clinical success and ultimately becoming an approved drug. Unfortunately, this step is also extremely expensive, time-consuming, and plagued with a high rate of failure. One of the biggest challenges is that lead optimization must simultaneously assure efficacy, safety, and developability. It is a very common case that perfecting one parameter ruins another – a process that has often been described as playing a game of ‘whack-a-mole’ where improving one property may lead to unintended effects on other properties and so on in a perpetual cycle that may never yield an ideal candidate.
Two of the most important properties to optimize – or moles to simultaneously knock down – before preclinical studies are efficacy and safety. These are indicators that an antibody will have a strong therapeutic effect and be less likely to have severe side effects. In order to have high efficacy a good therapeutic antibody candidate will ideally only attach, or bind, to very specific cells in your body, such as cancer cells. This allows the cancer cell to be destroyed by the immune system while leaving healthy surrounding tissue intact. The antibody binding must be not only specific, but also very strong and long-lasting to give immune cells time to kill the cancer cells. The strength of this binding is defined as affinity, and is a key metric drug developers seek to optimize for therapeutic antibodies.
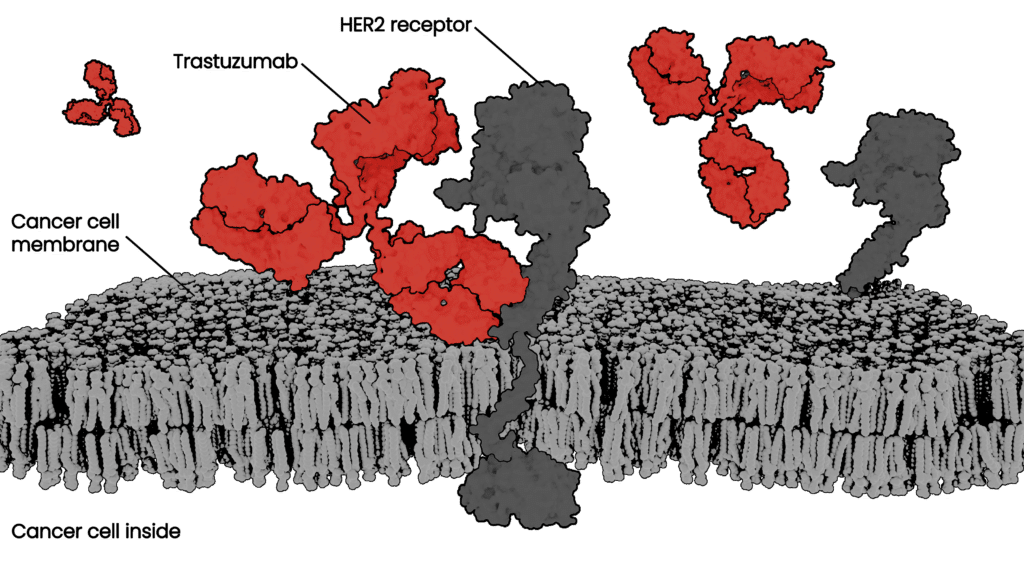
Another indicator of whether a therapeutic antibody will produce severe side effects is immunogenicity – or whether the antibody is likely to trigger an unwanted immune response. Immunogenicity in particular, can be a tricky parameter to gauge in a single experiment due to the complexity of the immune system and all the different cells and proteins a therapeutic antibody may encounter on the way to its target. However, it has been hypothesized that the more similar an antibody is to those occurring in nature, the less likely they will be to create an immune response. With this in mind, we have been developing a new metric called “naturalness”. We believe that therapeutic antibodies with high naturalness scores will be less likely to trigger an immune response in patients, leading to fewer adverse effects and better outcomes.
Stronger, Better, Faster: Machine Learning To The Rescue
To design a promising lead drug candidate for further development, you need to find antibodies with several different properties, all falling within a small range. It’s like searching for a needle in a haystack, except the haystack is larger than the known universe, and the needle is smaller than a speck of sand. Using traditional discovery methods, this can be a long, expensive process with a little probability of success. At Absci we’ve designed our generative AI models to significantly speed up and improve the lead optimization process compared to traditional methods that rely solely on wet lab screening. Using our proprietary wet lab assays we generate the massive amounts of biological data needed to enable the predictive power of machine learning. Using computers to design, test, and run experiments in silico (on the computer rather than at a lab bench) we can dramatically speed up the lead optimization process and discover unique, generative variants that could never be found using standard methods.
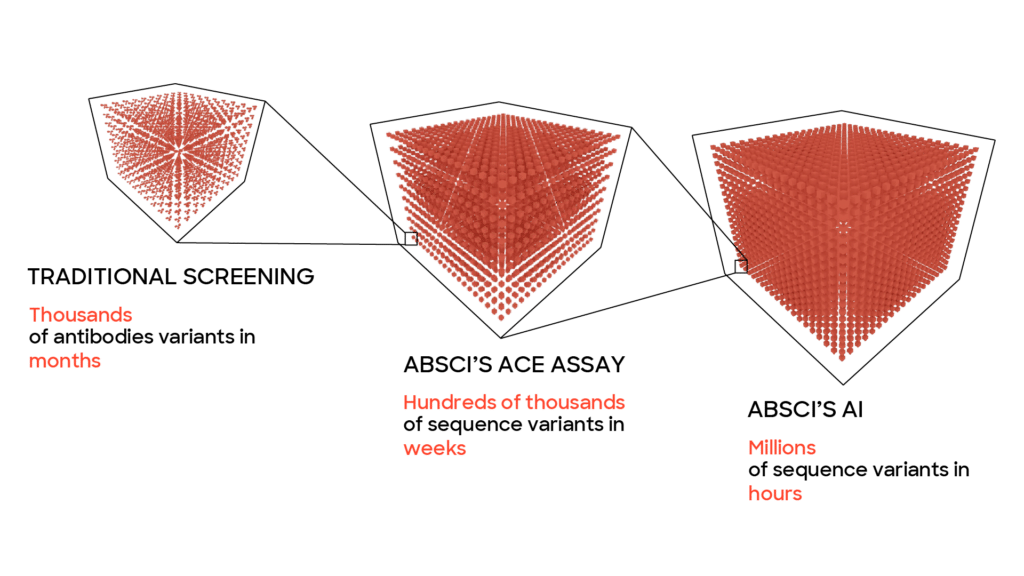
One reason this method is able to work faster than traditional methods is that machine learning models allow us to test a much larger number of potential drugs – far more than would be possible or practical to explore exclusively in a wet lab due to time and money considerations. With our platform, we can quickly probe a wide variety of vastly different potential drugs and then hone in on the most promising ones extremely quickly. To do what would take years using traditional methods can be done in months or weeks. For the lead optimization phase of a project this enables the computer to play our metaphorical whack-a-mole game at a breakneck pace until it finds an antibody where all the parameters have been optimized and the therapeutic antibody can continue to preclinical studies.
Let’s Work Together to Bring Therapeutic Antibodies to Patients Faster!
We work with our partners on target and antibody discovery, AI-assisted antibody optimization, and site-specific payload conjugation via nsAA incorporation. Connect with us today to see how we can help you create better biologics.